from triangulax.triangular import TriMeshMesh geometry
Using the half-edge mesh and the adjacency-like operators it defines, we can compute all sorts of geometric quantities of interest: edge lengths, cell areas, curvature in 3d, etc.
Discrete exterior calculus and discrete Hodge star
Note: triangle area, cell area, edge length, and dual edge lengths are what’s required for discrete exterior calculus.
# load test data
mesh = TriMesh.read_obj("test_meshes/disk.obj")
hemesh = msh.HeMesh.from_triangles(mesh.vertices.shape[0], mesh.faces)
geommesh = msh.GeomMesh(*hemesh.n_items, mesh.vertices, mesh.face_positions)
mesh_3d = TriMesh.read_obj("test_meshes/disk.obj", dim=3)
geommesh_3d = msh.GeomMesh(*hemesh.n_items, mesh_3d.vertices, mesh_3d.face_positions)Warning: readOBJ() ignored non-comment line 3:
o flat_tri_ecmc
Warning: readOBJ() ignored non-comment line 3:
o flat_tri_ecmcBasic mesh geometry
get_he_length
def get_he_length(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_hes']:
Get lengths of half-edges (triangulation/primal edges).
get_dihedral_angles
def get_dihedral_angles(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_hes ...']:
Get dihedral angles (angle between adjacent face normals).
get_vertex_normals
def get_vertex_normals(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_faces ...']:
Compute per-vertex normals by averaging over faces
get_triangle_normals
def get_triangle_normals(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_faces ...']:
Compute normals. In 2d, this just returns +/-1.
get_oriented_triangle_areas
def get_oriented_triangle_areas(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_faces ...']:
Compute oriented triangle areas in a mesh. In 3d, this is a vector.
get_triangle_areas
def get_triangle_areas(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_faces ...']:
Compute triangle areas in a mesh.
set_voronoi_face_positions
def set_voronoi_face_positions(
geommesh:GeomMesh, hemesh:HeMesh
)->GeomMesh:
Set face positions of geommesh to the circumcenters of the faces defined by hemesh.
get_voronoi_face_positions
def get_voronoi_face_positions(
vertices:Float[Array, 'n_vertices 2'], hemesh:HeMesh
)->Float[Array, 'n_faces 2']:
Get face positions of geommesh to the circumcenters of the faces defined by hemesh.
get_oriented_dual_he_length
def get_oriented_dual_he_length(
vertices:Float[Array, 'n_vertices 2'], face_positions:Float[Array, 'n_faces 2'], hemesh:HeMesh
)->Float[Array, 'n_hes']:
Compute lengths of dual edges. Boundary dual edges get length 1. Negative sign = flipped edge.
get_dual_he_length
def get_dual_he_length(
face_positions:Float[Array, 'n_faces dim'], hemesh:HeMesh
)->Float[Array, 'n_hes']:
Get lengths of dual/cell half-edges.
a = get_dual_he_length(mesh.face_positions, hemesh)
b = get_oriented_dual_he_length(mesh.vertices, mesh.face_positions, hemesh)
jnp.allclose(a[~hemesh.is_bdry_edge], jnp.abs(b)[~hemesh.is_bdry_edge])Array(True, dtype=bool)# edges and dual edges should be orthogonal since we are using circumcenters
face_positions = get_voronoi_face_positions(mesh.vertices, hemesh)
edges = mesh.vertices[hemesh.orig]-mesh.vertices[hemesh.dest]
dual_edges = (face_positions[hemesh.heface]-face_positions[hemesh.heface[hemesh.twin]])
jnp.allclose(jnp.einsum('vi,vi->v', edges[~hemesh.is_bdry_edge], dual_edges[~hemesh.is_bdry_edge]), 0)Array(True, dtype=bool)# computing the signed edge length shows that there are some "flipped" edges.
dual_length = get_oriented_dual_he_length(mesh.vertices, face_positions, hemesh)
jnp.where((dual_length < -0.0) & ~hemesh.is_bdry_edge )[0]Array([ 9, 185, 191, 335, 363, 539, 545, 689], dtype=int64)get_voronoi_edge_lengths
def get_voronoi_edge_lengths(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_hes']:
Computed directly from angles. Accurate in any dimension
get_cotan_weights_per_egde
def get_cotan_weights_per_egde(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_hes']:
Average of cotangent of angles opposite to edge.
get_cotan_weights_per_he
def get_cotan_weights_per_he(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_hes']:
Cotangent of angle opposite to half-edge
get_angle_sum
def get_angle_sum(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_vertices']:
Angle sum around vertices. 2pi-angle sum measures Gaussian curvature*
get_corner_angles
def get_corner_angles(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_hes']:
Get angles in mesh corners (opposite to half-edges).
angles = get_corner_angles(mesh.vertices, hemesh)
np.allclose(get_angle_sum(mesh.vertices, hemesh)[~hemesh.is_bdry], 2*jnp.pi) # mesh is not curvedTruejnp.allclose(1/jnp.tan(angles), get_cotan_weights_per_he(mesh.vertices, hemesh))Array(True, dtype=bool)# we can either compute the Voronoi-length of a dual edge directly, or from the face positions
voronoi_edge_lengths = get_voronoi_edge_lengths(mesh.vertices, hemesh)
dual_edge_length = get_oriented_dual_he_length(mesh.vertices, mesh.face_positions, hemesh)jnp.allclose(voronoi_edge_lengths[~hemesh.is_bdry_edge], dual_edge_length[~hemesh.is_bdry_edge])Array(True, dtype=bool)Computing cell areas, perimeters, etc via corners
To compute, for instance, the cell area using the shoelace formula, you need to iterate around the faces adjacent to a vertex. This is not straightforward to vectorize because the number of adjacent faces per vertex can vary (there can be 5-, 6-, 7-sided cells etc.). One way to solve this is a scheme in which the lists of adjacent faces are “padded” in some manner, so that they are all the same length. This is cumbersome. Instead, let us split all “cell-based” quantities into contributions from “corners”, i.e., half-edges, like this:
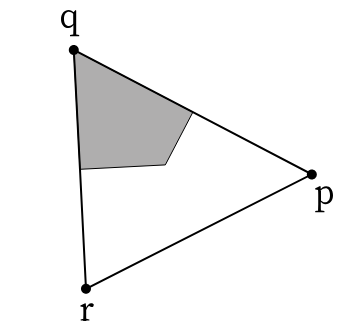 Source: CGAL
Source: CGAL
To compute the total area, we can sum over all half-edges \((r,p)\) opposite to a vertex \(q\).
This approach can also compute cell perimeter, …
get_voronoi_areas
def get_voronoi_areas(
vertices:Float[Array, 'n_vertices dim'], hemesh:HeMesh
)->Float[Array, 'n_vertices']:
Compute Voronoi area for each vertex.
get_cell_areas_traversal
def get_cell_areas_traversal(
geommesh:GeomMesh, hemesh:HeMesh
)->Float[Array, 'n_vertices']:
Compute areas of cells by mesh traversal (don’t use for simulation, inefficient).
Boundary vertices get area 0.
## Let's use the adjacency matrix to compute the area of all cells. First, compute all corner areas
a, b, c = (hemesh.dest[hemesh.nxt], hemesh.dest[hemesh.prv], hemesh.dest)
corner_areas = jax.vmap(trig.get_voronoi_corner_area)(geommesh.vertices[a], geommesh.vertices[b], geommesh.vertices[c])
cell_areas_corner = adj.sum_he_to_vertex_opposite(hemesh, corner_areas)
cell_areas_corner = cell_areas_corner.at[hemesh.is_bdry].set(0)# for comparison, compute the areas by mesh traversal
cell_areas_iterative = get_cell_areas_traversal(geommesh, hemesh)
np.abs(cell_areas_iterative-cell_areas_corner).max() # works!np.float64(5.551115123125783e-17)